T84-IL22-IFNL
T84 cells treated with IFNL and IL22
Experimental design
T84 cells were treated with the cytokines, type III interferon lambda (IFN-L), interleukin 22 (IL-22), and in combination (IFN-L + IL-22) at time points - 3h, 6h, 12h and 24h. Each time point and treatment were replicated three times. RNA sequencing was performed in a total of 48 samples consisting of untreated and treated samples.
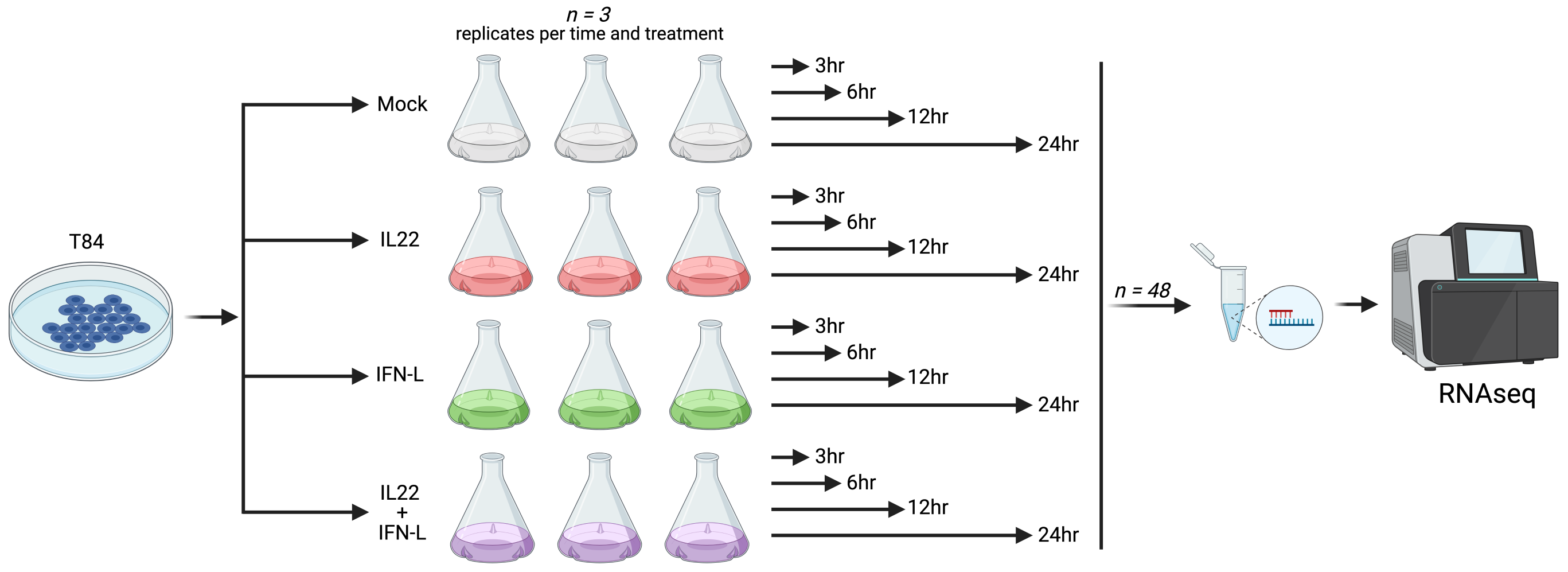
Methodology
Raw RNA sequencing reads (fastq) were aligned to the Ensembl human GRCh38 genome reference using Rsubread with default settings. Read summarization was done using featureCounts. Various quality metrics of the raw reads and alignment statistics were analyzed using MultiQC. Differential gene expression analysis was performed using DSeq2, rlog transformed data was used for multi-dimensional scaling (MDS) and unsupervised hierarchical clustering analyses. Signaling programs were quantified using PROGENy. Transcription factor activities were computed using DoRothEA and VIPER. Enrichment analysis on the most differentially expressed genes (-1< logFC >+1 and adjusted p-value < 0.05) was performed using enrichR.
Data download
Raw and processsed data from this study can be downloaded below. The source code for this study is available here
- Raw fastq files
- Raw counts - RDS, csv
- rlog transformed data - RDS
- Quality control - MultiQC
- Differentially expressed genes (All) - RDS, xls
- Differentially expressed genes - IL22
- Differentially expressed genes - INFL
- Differentially expressed genes - IL22 + INFL
- log2 fold changes (RDS)
- Signalling programs
- Transcription factor activities